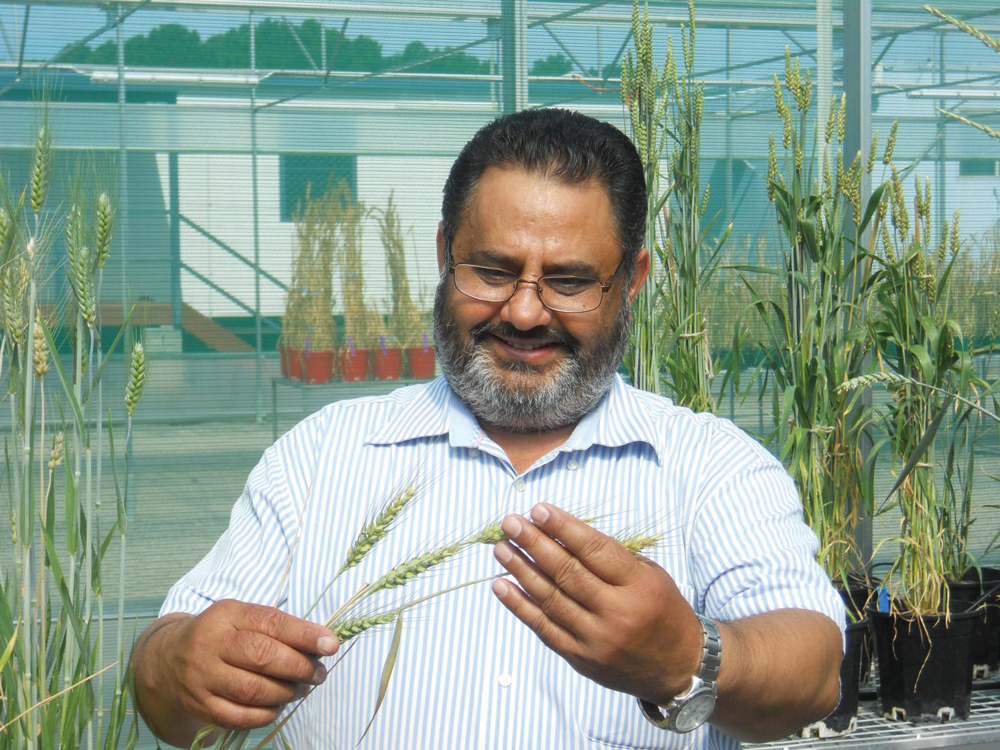
A global alliance of researchers has pioneered a new method to rapidly recruit disease-resistance genes from wild plants for transfer into domestic crops.
The technique promises to revolutionize the development of disease-resistant varieties.
The technique called AgRenSeq was developed by scientists at the John Innes Centre in Britain working with colleagues in Australia and the U.S.
The result speeds up the fight against pathogens that threaten global food crops, including wheat, soybean, maize, rice and potato.
Harbans Bariana from the Sydney Institute of Agriculture and the School of Life and Environmental Sciences is a global expert in cereal rust genetics and a co-author of the paper.
Read Also

Indo-Pacific seen as potential organics market
Organic producers who want to export Canadian agrifoods should look to the Indo-Pacific region, an expert on trade in the area says.
“This technology will underpin fast-tracked discovery and characterization of new sources of disease resistance in plants,” Bariana said.
The current research builds on previous collaborative work and used two wheat genes cloned by this international team as controls.
AgRenSeq lets researchers search a library of resistance genes discovered in wild relatives of modern crops so they can rapidly identify sequences associated with disease-fighting capability.
From there, researchers can use laboratory techniques to clone the genes and introduce them into elite varieties of domestic crops to protect them against pathogens and pests such as rusts, powdery mildew and Hessian fly.
Brande Wulff, a crop genetics project leader at the John Innes Centre and a lead author of the study, said: “We have found a way to scan the genome of a wild relative of a crop plant and pick out the resistance genes we need.”